
This activity gives AP students the opportunity to work with the protein database: UniProt. This tool can align protein sequence and perform BLAST.
Once students are familiar with the tool
Students become familiar with the tool by answering the question “Are Bats Birds” by comparing the protein sequence of hemoglobin between birds, bats, and other mammals. Hemoglobin will show greater similarities between bats and other mammals than with bats and birds. Each comparison provides a percent identity for how many of the amino acids are the same. The higher the percent, the greater the relationship. The tool also has a function where a phylogenetic tree can be created based on those similarities.
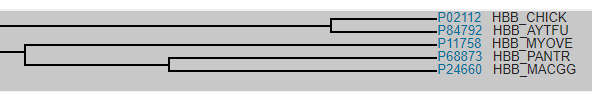
Once students are familiar with the tool, they are tasked with determining what organisms would make good models for studying the Cystic Fibrosis Transmembrane Protein (CFTR). There are less instructions, but students should be able to develop a chart of phylogenetic tree that shows what animals have similar CFTR proteins.
An optional investigations asks them to look at whales and their relationship to land animals . This one I usually give as an optional task either for extra credit.

