Restiction Enzymes: How is DNA Manipulated?
Using restriction enzymes, foreign genes can be added to an existing organism (or an embryo). This organism has been genetically modified. Adding new genes can create plants that are more resistant to pests or be more tolerant to weather patterns, such as drought. This technology can also be used to mass produce chemicals, such as human growth hormone, by inserting that gene into bacteria.
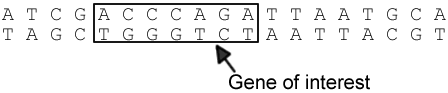
In order to combine the DNA, a chemical called a restriction enzyme is used to cut the DNA into fragments, exposing the gene of interest. On either side of the gene is an area of DNA called the “sticky end.” The bases of the sticky end are ready to be paired to the new DNA following the base-pair rule. That gene is then spliced to the other organism’s DNA using another enzyme called ligase, the joining procedure is called ligation.
Sometimes scientists want a gene removed without the sticky ends, so that it will not bind to other parts of DNA. These fragments are said to have “blunt ends”.
*Recombinant DNA is often abbreviated as rDNA to denote that it has foreign genes (DNA) inserted into its genome.
This image shows a restriction enzyme called EcoRI being used to cleave a section of DNA. Different restriction enzyme will cleave DNA at different sequence points, called recognition sites, so scientists must isolate the right enzyme for the job – one that cuts around the desired gene.
You will often see restriction enzyme sites written like this: G / A A T T C.
How is DNA Manipulated?
1. Explain the following terms and their role in recombinant DNA technology
a) Restriction enzyme _____________________________________________________________b) Recognition site ________________________________________________________________
c) Sticky end ____________________________________________________________________
d) Ligation ______________________________________________________________________
e) rDNA ______________________________________________________________________
Here are some restriction enzymes and their sites.
| BAMHI | G / G A T C C | PstI | C T G C A / G |
| HindIII | A / A G C T T | HhaI | G C G / C |
| EcoRI | G / A A T T C | HpaII | C / C G G |
| SalI | G / T C G A C | ||
| HindII | G T C / G A C C A G / C T G (blunt ends) |
||
2. On each of the sequences below, determine which restriction enzyme could be used to splice the DNA and indicate where the cut will be made and the enzyme used.
| 1. | 5' T T T G A A T T C A G A T 3' 3' A A A C T T A A G T C T A 5' |
Enzyme: |
| 2. | 5' G T G G G A T C C C T T A 3' 3' C A C C C T A G G G A A T 5' |
Enzyme: |
| 3. | 5' A C G C C T C C G G A G A 3' 3' T G C G G A G G C C T C T 5' |
Enzyme: |
| 4. | 5' T T A A G C T T A A G A A G C T T 3' 3' A A T T C G A A T T C T T C G A A 5' |
Enzyme: |
| 5. | 5' A A G C G C G T C G A C T T A T A 3' 3' T T C G C G C A G C T G A A T A T 5' |
Enzyme: |
3. Create a restriction enzyme that will remove the gene of interst. Give it a name too!
4. The following DNA sequence is from a virus that is dangerous, scientists want to use a restriction enzyme to cut the virus into bits. They do not need sticky ends because the do not plan to combine it with other DNA. Use HindII to show how this DNA would be cut. How many pieces would you have? ____
C T T T T C A G C T G T T C C G T C A G C T G A A A A T T T T C A G C T G T A C G
How is the Recombinant DNA (rDNA) Used?
a) Which genes are being isolated from the flu virus?
b) Where are the flu genes (HA and NA) placed?
c) Where do plasmids come from and why are they important in this process?
d) What are the potential applications for making a new flu?
Related Resources
DNA Crossword - puzzle to practice basic terms related to DNA
DNA Detectives: The Case of the Radioactive Phosphorous - a short story about the Hershey Chase experiment with questions.
Regulatory Switches in the Stickleback - based on an HHMI activity, explores gene expression in fish (Also available as a slide activity)
Gene Regulation in Prokaryotes - focuses on the lac operon and how some genes are repressible