Drosophila Simulation - Patterns of Heredity
Objective: Students will learn and apply the principles of Mendelian inheritance by experimentation with the fruit fly Drosophila melanogaster. Students will make hypotheses for monohybrid, dihybrid and sex-linked traits and test their hypotheses by selecting fruit flies with different visible mutations, mating them, and analyzing the phenotypic ratios of the offspring.
Website: https://www.sciencecourseware.org/FlyLabJS/
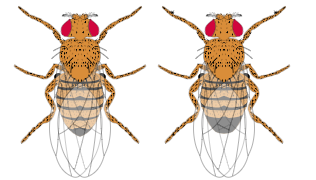
The image shown below shows a wild-type female fly (left) and a male fly. Recall that "wild-type" refers to the most common or typical form seen in the wild. A + sign is used to denote when a fly displays the wild-type characteristic.
Introduction
Examine the phenotypes available from the left side menu to answer the following questions.
1. Examine the different types of bristles seen in flies. Geneticists use a shorthand labeling system, F = forked. Identify the phenotypes shown:


2. Compare antennae types. How is "aristapedia" different from wild-type?
3. What are different eye colors in fruit flies? Circle the one that is wild-type.
4. Regarding wing size, what is the difference between apterous and vestigial?
5. What are the body colors in fruit flies?
6. Create a mutant fly with any number of variations and mate it with a wild-type fly. How many offspring were wild-type?
7. Mate the offspring of the cross. Use the analyze tab to get more details about the F2 offspring. (The button to "ignore sex" may make counting easier.)
How many wild-type offspring were produced?
How many mutant flies were produced?
Part 2: Monohybrid Crosses
You may realize that choosing a lot of different types of flies make it difficult to analyze inheritance patterns. Your next tasks will focus on analyzing single traits within flies to determine how they are inherited.
1. Reset all flies in the design tab.
2. Design a male fly with vestigial wings and cross it with a wild-type female
3.
Add the results to your "Lab Notes."
4.
Mate the offspring of this cross.
5. Based on these two crosses you probably have an idea about how vestigial wings are inherited.
Is VG recessive or dominant?
How do you know?
6. In genetics, numbers are statistically analyzed. The fly simulator has a built into it. Under the Analyze tab, you can click on "Include a test hypothesis."
If your hypothesis that VG is a recessive trait is correct, then you would expect what proportion of the F2 offspring to have vestigial wings?
What proportion would have wild-type wings?
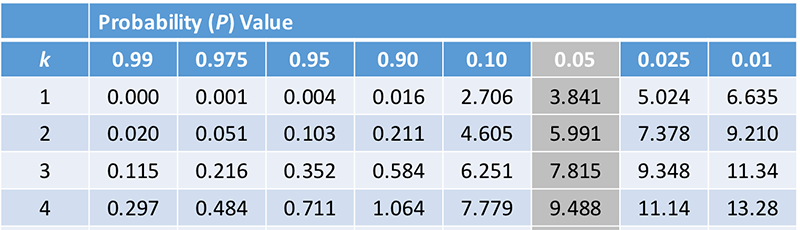
7. Place the expected numbers in the hypothesis field and click on "test your hypothesis." The program will do the chi square calculations.
What is your chi-squared test statistic?
Compare this to the chi square table to determine a goodness of fit.
8. Summary: Explain how vestigial wings are inherited in fruit flies (claim) and provide evidence from your data and chi-square statistic analysis.
Part 3: Sex Linked Traits
1. Cross a white eyed male with a wild-type female.
How many of the offspring are males / red eyes?
How many females / red eyes?
2. Predict what would happen if you crossed two of the offspring. Explain your reasoning by showing a punnett square
3. Perform the cross and use the statistical analysis tool to test your prediction.
4. Summary: Explain how red/white eye color is inherited in fruit flies (claim) and provide evidence from your data and chi-square statistic
Part 4: Lethal Alleles
Aristapedia is a lethal allele that is also dominant. Individuals with this trait must be heterozygous (Aa) because the homozygous condition (AA) is lethal. This is not a sex-linked trait. Wild-type flies do not carry the allele for aristopedia (aa).
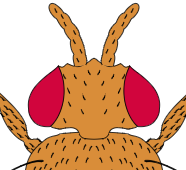

1. Predict what the outcome of a cross between a wild-type fly and one with aristopedia. Show the punnett square to illustrate your reasoning.
2. Perform the cross and determine if your prediction is correct using statistical analysis. Summarize your results and indicate whether your prediction is confirmed.
Part 5: Linkage Groups
When two alleles are located on the same chromosome they are inherited together. However, crossing-over can occur during meiosis and the alleles are switched. Vestigial wings (VG) and Black body color (BL) are located on chromosome 2.
1. Cross a female VG, BL fly with a wild-type male. (ggbb x GGBB)
How many wild-type offspring are produced?
What is the genotype of these offspring?
2. Choose a female from the offspring and mate it with a male that has vestigial wings and a black body. Show a punnett square or a visual representation of the alleles involved in this cross to make a prediction about the offspring.
3. Complete the table (ignore sex).
| Phenotype | Observed | Proportion |
| + (wild-type) | ||
| Vestigial wings (gg) | ||
| Black body (bb) | ||
| VG, BL (ggbb) |
4. How does crossing-over affect the observed outcomes? Explain why the observed flies do not match your prediction.
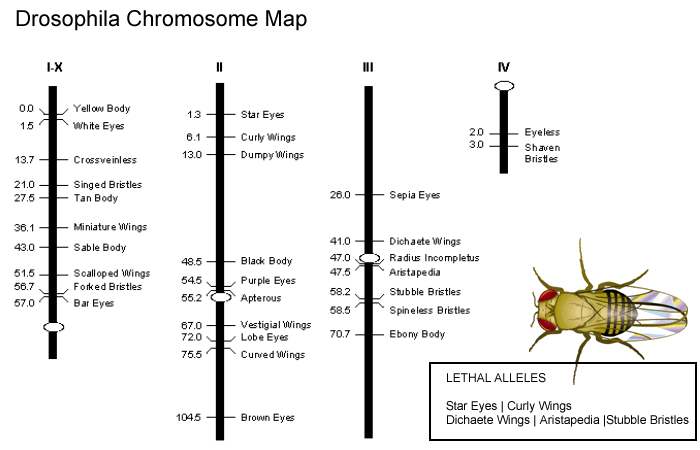
5. The percentage of crossing over events is used to develop a map of chromosomes. View the chromosome map.
How far apart are the alleles for black body and vestigial wings?
View the proportion of flies from your data that indicate crossover occurred (VG and BL flies) and multiple it by 100. Based on your data, how far apart are these alleles?
Resources
Other Resources on Genetics
Corn Genetics and Chi Square - statistical analysis, using preserved corn and counting kernels
Genetics of Wisconsin Fast Plants - grow plants and analyze phenotype ratios
Albino Corn Genetics - grow corn, 3:1 albino ratio, lab report analyzes F1, F2 crosses